今天分享一个自己写的python小脚本可以实现单碱基的基因组位置转换到转录本的坐标,欢迎大家使用,并提出错误
#!/share/home/hujun/miniconda3/bin/python3 import pybedtools from pybedtools import BedTool import pandas as pd import argparse def intersect_region(genepred_file, genome_site_file): genepred = pd.read_csv(genepred_file, sep="\t", header=None) genepred = genepred.iloc[:, [1, 3, 4, 0, 7, 2, 5, 6, 8, 9]] genepred = BedTool.from_dataframe(genepred) genome_site = pybedtools.BedTool(genome_site_file) intersect = genepred.intersect(genome_site, wa=True, wb=True, s=True) return intersect def transform_position(intersect): output = [] for i in intersect: site = int(i[12]) exon_start = i[8].strip(",").split(",") exon_end = i[9].strip(",").split(",") exon_number = i[4] id = i[3] strand = i[5] if strand == "+": length = 0 for j in range(len(exon_start) - 1): length = length + int(exon_end[j]) - int(exon_start[j]) if site >= int(exon_start[j]) and site <= int(exon_end[j]): position = length - (int(exon_end[j]) - site) tras_p = [id, position] output.append(tras_p) else: length = 0 for j in range(len(exon_start) - 1, 0, -1): length = length + int(exon_end[j]) - int(exon_start[j]) if site >= int(exon_start[j]) and site <= int(exon_end[j]): position = length - (site - int(exon_start[j])) + 1 tras_p = [id, position] output.append(tras_p) return pd.DataFrame(output) def id_change(): intersect = intersect_region(genepred_file=genepred, genome_site_file=bed) out = transform_position(intersect) trascript_index = pd.read_csv(index, sep="\t", engine="python", header=None) new_col = trascript_index.iloc[:, 0].str.split("|", expand=True) trascript_index["id"] = new_col[0] a = pd.merge(out, trascript_index, how="left", left_on=0, right_on="id") outdf = a.iloc[:, [3, 2, 0]] outdf.to_csv(output, sep="\t", index=False) if __name__=="__main__": parser = argparse.ArgumentParser( description='This script aim to transform a single genome position to the transcript position', add_help=True) parser.add_argument('-g', '--genepred', type=str,required=True, help='The genepred format file which cat transform from the gtf file') parser.add_argument('-b', '--bed', type=str,required=True, help='The bed format file for a single position of the genome ,such as SNP,' 'and must have six columns', ) parser.add_argument('-d', '--index', type=str,required=True, help='The transcript fasta index file which you can generate with the ' 'samtools') parser.add_argument('-o', '--output', type=str, required=True,help='The output filename') args = parser.parse_args() if not any(vars(args).values()): parser.print_help() else: genepred = args.genepred bed = args.bed index = args.index output = args.output id_change()
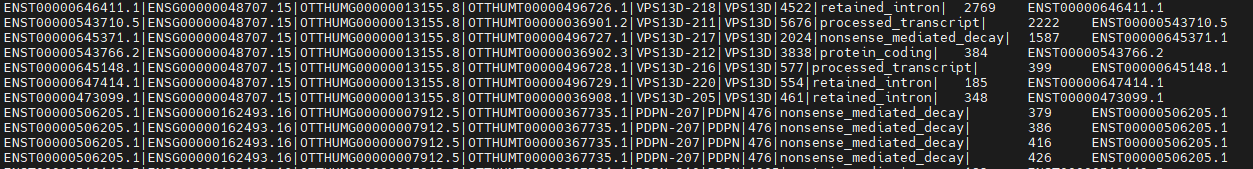
最终输出的文件总共有三列,第一列为转录本fasta文件中的名字,第二列为1 base 的转录本坐标,第三列为转录本id,之所以设置成这个样子是为了方便和一些比对到转录本中的数据进行比较