001、使用命令 及生成结果
samtools flagstat -@ 56 ERR3143219.sort.bam > flagstat.txt ## samtools flagstat统计
002、输出结果解读
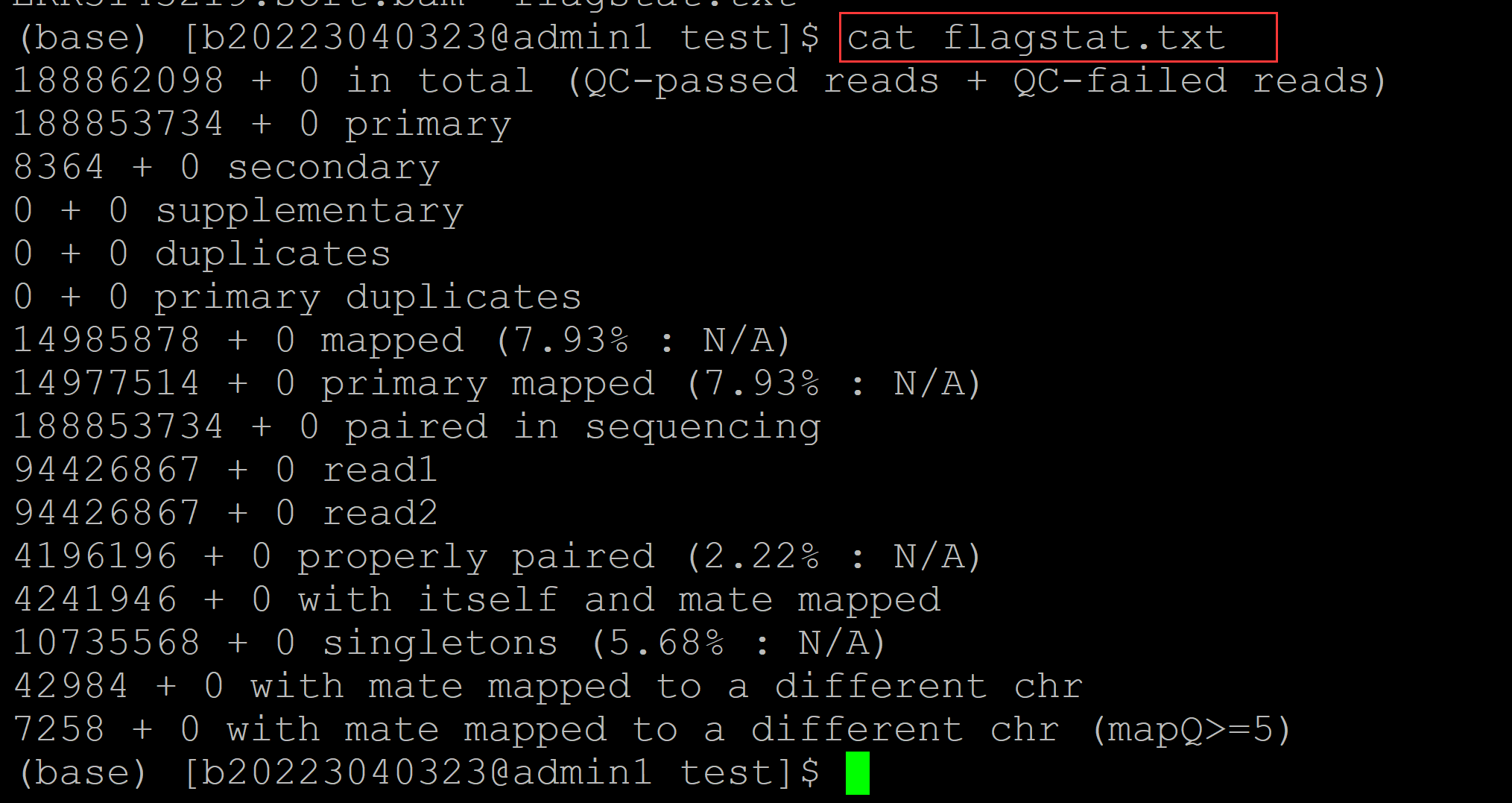
(base) [b20223040323@admin1 test]$ cat flagstat.txt ## 会多于原始的fastq中的reads数目,因为存在比对到参考基因组多个位置的情况 188862098 + 0 in total (QC-passed reads + QC-failed reads) ## 比对到参考基因组总的read数目,包括原始的fastq的总的reads数和和比对到参考基因组多个位置reads的计数 188853734 + 0 primary ## 原始的fastq中的read数目 8364 + 0 secondary 0 + 0 supplementary 0 + 0 duplicates 0 + 0 primary duplicates 14985878 + 0 mapped (7.93% : N/A) 14977514 + 0 primary mapped (7.93% : N/A) 188853734 + 0 paired in sequencing 94426867 + 0 read1 94426867 + 0 read2 4196196 + 0 properly paired (2.22% : N/A) 4241946 + 0 with itself and mate mapped 10735568 + 0 singletons (5.68% : N/A) 42984 + 0 with mate mapped to a different chr 7258 + 0 with mate mapped to a different chr (mapQ>=5)
。
参考:
01、https://wenku.baidu.com/view/94ef844924d3240c844769eae009581b6bd9bd96.html?_wkts_=1699923563989&bdQuery=samtools+flagstat+%E5%8F%82%E6%95%B0+%E7%BB%93%E6%9E%9C%E8%A7%A3%E8%AF%BB